Research
The liverwort way of being a plant
Liverworts diverged from the rest of landplants more than 400 My ago. Since then, they have evolved unique structures, genetic pathways, and life history strategies among land plants. In comparison to the most abundant and comparatively better studied vascular plants, liverworts are small. The dominant phase of these plants is the gametophytic haploid fase, which displays a wonderful diversity of morphologies and life strategies that are not observed in any other lineage.
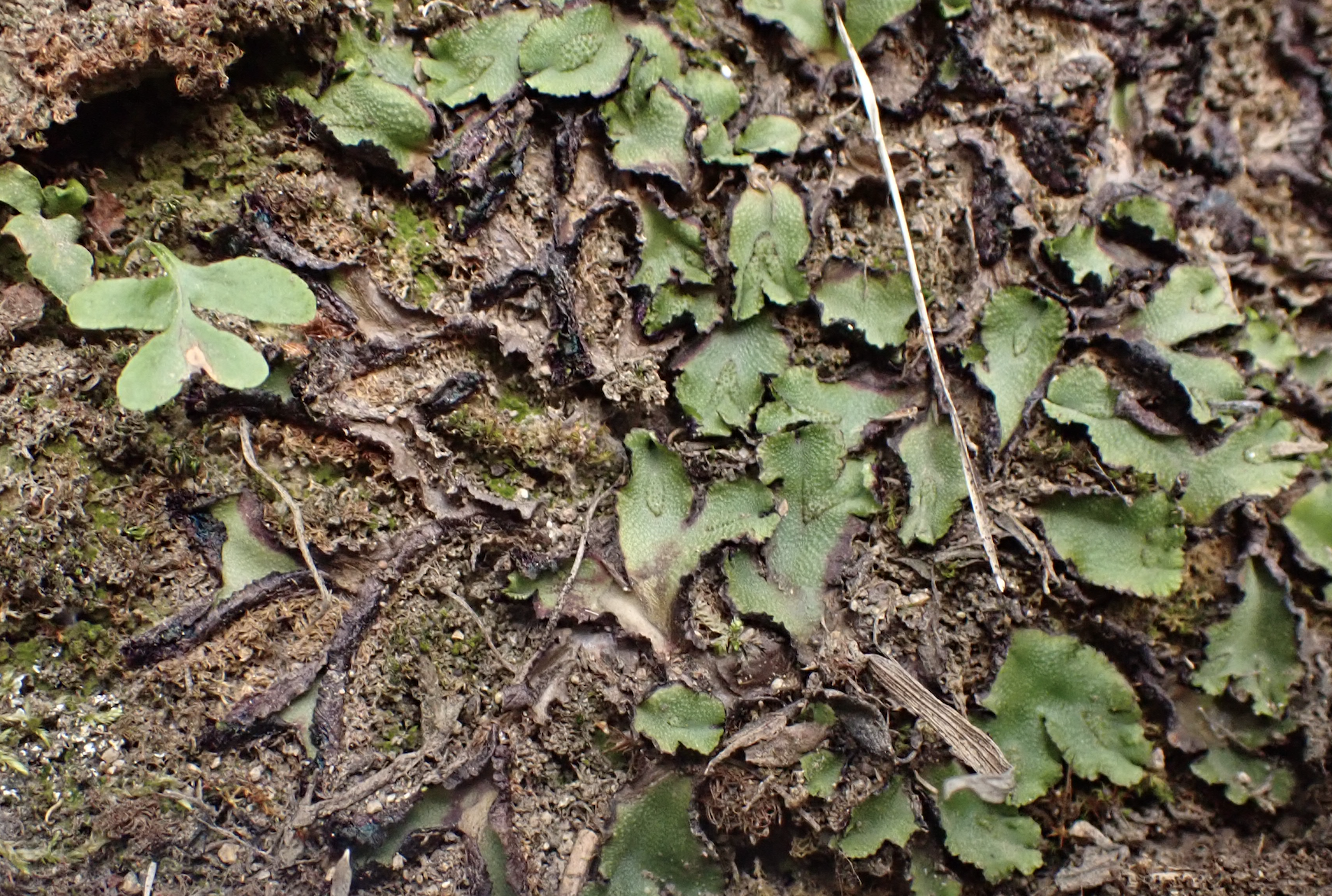

Liverwort gametophytes come in multiple forms, from thallose to leafy. Sporophyte bearing structures are also extremely diverse, from incospicuous sacs embeded in the thalli to showy umbrella-like archegoniophores. They have unique organelles (oil bodies) and structures (e.g., ventral scales, pegged rizoids, elaters, archegoniophores, antheridiophores). They are known to have strong interactions with fungi, and (recently argued) bacteria.
One of the main curiosity drivers of this lab, is to explore the liverwort way of being a land plant. What strategies do these plants have evolved to thrive in diverse environments? What genetic, environmental, and developmental constraints do they fase? How unique is the liverwort way?
While thinking about unique structures in liverworts, we have focused on the ventral scales–dermal structures that are very diverse across taxa. Our preliminary research has shown that purple ventral scales can absorbe UV radiation, so it is no surprise that xerophytic liverworts like Targiona hypophylla and Calasterella californica display a “roll up” mechanism, in which the dessicated thalli wrap themselves with their ventral scales, which potentially provide UV protection during a vulnerable physiological state.
A couple lines of research on the liverowort way in the lab are (1) exploring the morfological and functional diversity of ventral scales in complex thalloid liverworts, and (2) characterizing bacterial communities associated with liverworts.
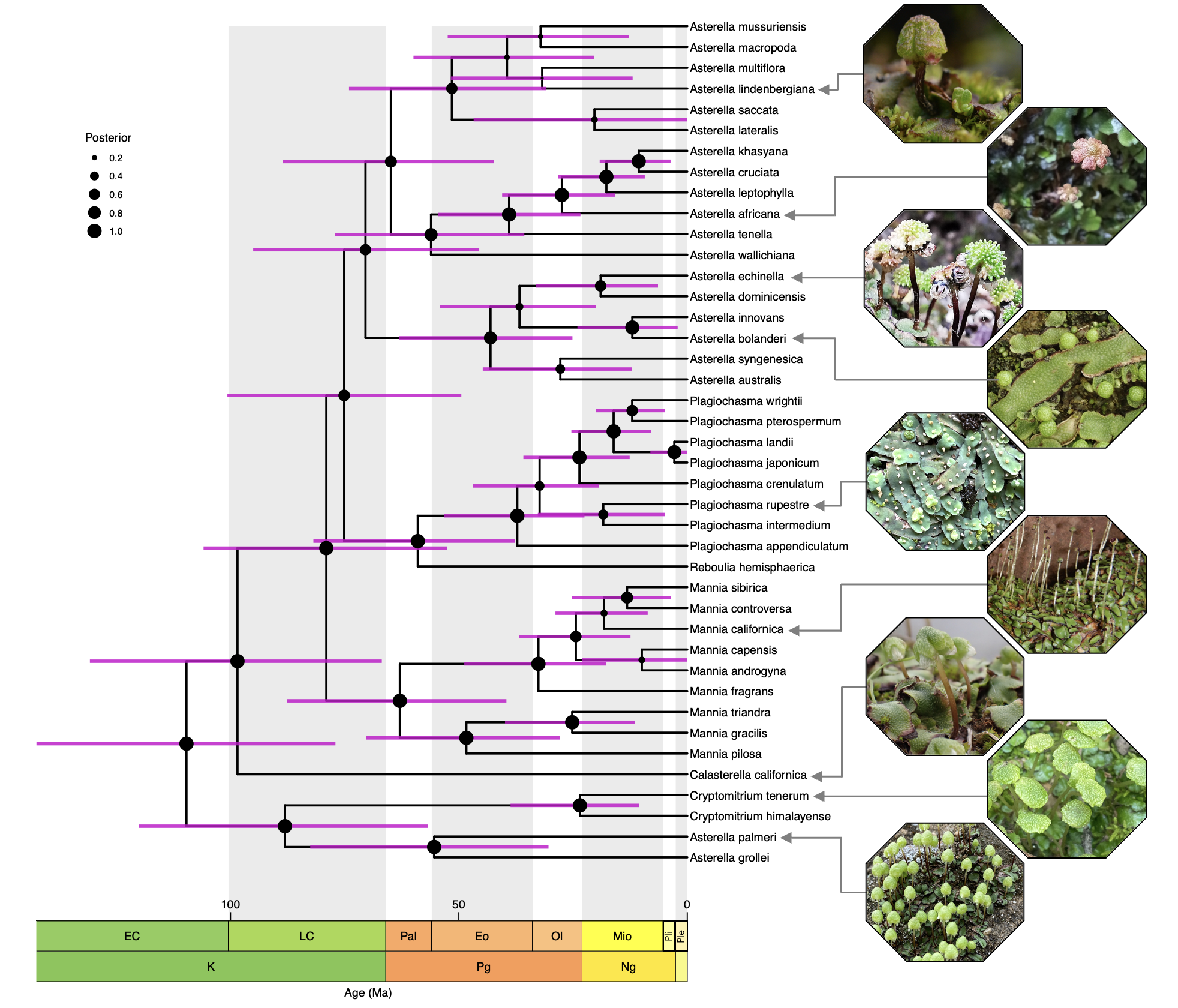
Evolutionary history of complex thalloid liverworts
We strive to infer patterns and understand the processes that have driven liverwort evolution. How does their unique biology is the result of evolutionary processes? What determined their distribution? Are evolutionary drivers of liverwort evolution the same as in other groups? Why do they have such broad distributions? Some of our ongoing projects on this regard are focused on the biogeographical history of Aytoniaceae, a family of complex thalloid liverworts, and the effect that spore morphology might have on the regions that different species occupy, e.g., are UV protective scales a prerequisite for a xerophytic habit.
Integrative models of evolution
Our prefered way to address all these evolutionary questions is to use model evolutionary processes that account for the complexity of biological processes. These models (e.g., FBD, SSE family of models) often co-estimate multiple parameters of interest simultaneously (trees, trait evolution, diversification events), and provide a natural framework to account for uncertainty—we believe it is important to have a sense of what we can’t know. An additional feature of these models is that they can often use multiple sources of information (e.g. geological, biological of different kinds) as they model multiple interacting features (e.g. how landmasses proximity affect lineage evolution). The implementation of these models is as challenging as it is rewarding, and it is an enterpise in itself. In collaboration with model developers, we implement exciting new models that fit our biological questions. Similarly, we love to collaborate with other researchers to implement some of our favorite models in groups with exciting datasets.
Information for prospective students:
The best fit for my lab would be students that are interested in either liverwort biology and evolution, or in integrative phylogenetic models such as the ones described above, or both! Having a background in systematics, phylogenetics, and basic coding skills would be welcome especially for graduate students, but it won’t be a limitation if the student is otherwise a good fit! I value curiosity and communication. If you have questions, reach out to me!
